Difference between revisions of "MBV-INFX410 2013"
Jonkl@uio.no (talk | contribs) |
Jonkl@uio.no (talk | contribs) |
||
| Line 333: | Line 333: | ||
<span id="3DChrom">On Wednesday November 20, between 11:15 and 12:00, there will be a combined course lecture and CLS seminar by Jonas Paulsen from Oslo University Hospital. This will be on the study of [http://nar.oxfordjournals.org/content/41/10/5164.abstract chromatin 3D structure]. </span> | <span id="3DChrom">On Wednesday November 20, between 11:15 and 12:00, there will be a combined course lecture and CLS seminar by Jonas Paulsen from Oslo University Hospital. This will be on the study of [http://nar.oxfordjournals.org/content/41/10/5164.abstract chromatin 3D structure]. </span> | ||
| − | === Exam === | + | === Exam === |
| − | + | The exam for this course will be a one week, take-home exam. | |
| + | |||
| + | The exam will be sent to all participants at 3 pm, Thursday December 6, by e-mail. | ||
| + | |||
| + | Your completed exam must be returned, at the latest, at 3 pm, Friday December 14. It should be sent by e-mail to the course administrator Torill Rørtveit (e-mail address: torill.rortveit@ibv.uio.no). Please put the course code and your candidate number in the subject field (''e.g.'' "Exam MBV-INF4410 Candidate:12345"). | ||
| + | |||
| + | The exam must be handed in as a single PDF document (Microsoft Word or an Open Office Document is also acceptable). The document should be named with the course code and your candidate number only (''e.g.'' MBV-INF4410-12345.pdf). '''''Do not place your name in the document.''''' | ||
=== Written assignment === | === Written assignment === | ||
Revision as of 17:12, 27 October 2013
Contents
Bioinformatics for Molecular Biology - 2013
Please bookmark this page. All future changes or announcements for the 2013 course will be posted on this page.
This is the wiki for the courses MBV-INF4410, MBV-INF9410, and MBV-INF9410A offered by the Department of Biosciences and Department of Informatics at the University of Oslo (UiO).
The course consists of five weeks of lectures, exercises, a take-home exam (one week), and a take-home exam/assignment (roughly 20 pages). The course is open also for non-UiO students. It is only necessary to be physically present in Oslo for certain parts of the course.
Ph.D. level students may opt to take the course without the take-home partial exam/assignment for only 8 study points (MBV-INF9410A). Both MBV-INF4410 (M.Sc. level) and MBV-INF9410 (Ph.D. level) are 10 study point courses.
Course description
This intensive course will introduce students to bioinformatics resources and tools for molecular biology research. All the lecturers are among the top researchers working within the fields of bioinformatics and computational biology in the Oslo region. Students must bring their own lap-top for in-course demonstrations as well as for practical lab exercises. The course is mainly intended for biology students, but also for computer science students or students from other fields of science with an interest for and some experience with molecular biology. No prior background in bioinformatics or computer science is required. All students should have a basic understanding of molecular biology, at least roughly corresponding to 5-10 university study points in molecular biology, biochemistry, or similar. If you are uncertain if your biology background is strong enough, please contact Jon (See contact details below) before you sign up for the course.
Course responsible is Dr. Jon K. Lærdahl (jonkl@medisin.uio.no) from the Department of Microbiology, Oslo University Hospital (OUH) - Rikshospitalet. Lærdahl is also employed by the CLS initiative at UiO and the Bioinformatics Core Facility (CF) at OUH and UiO.
Links to the web pages for the years 2009-2011 is found here and for 2012 here.
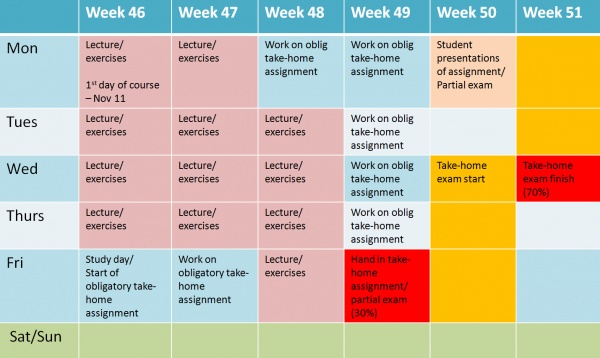
Notes on the course format: The course has previously been given as an intensive course over two weeks with a take-home exam in the 3rd to 4th week. A take-home assignment was also a compulsory part of the 10 study points versions of the course. The compact format was ideal for students coming from outside Oslo, but it was also exhausting for students and lecturers. It gave little time to digest and dive more deeply into the various topics presented in the course. In 2013, the course will be given in weeks 46-51. However, it will only be necessary to be physically present in Oslo for parts of the course, i.e. the lectures/exercises and, if practically possible, the students presentation day. The schedule is presented below, but there might be small adjustments to this schedule. The course content will be similar to the course in 2012.
Time and place
The course will be offered in weeks 46 to 50, autumn 2013, i.e. starting on Monday November 11 (See schedule below). The take-home exam must be handed in in week 51, on Wednesday December 18. Each day of lectures/exercises will consist of three time slots for lectures and/or exercises/practical labs between 09:00 and 16:00. Lunch will usually be between 12:45 and 13:30. You will have to bring your own lunch or buy lunch in the local kantine.
Lecture room: All lectures/exercises in weeks 46 and 47 (Monday to Thursday) will be given in lecture room Python in Ole-Johan Dahls hus (IFI2). A map showing the location of the building is found here. The building is located next to the Forskningsparken metro and tram stations. The room Python is on the 1st floor (2. etasje) in the northern end of the building, the end closest to the tram line. The easiest access to Python is through the entrance in the tunnel going through the building.
Lectures/exercises in week 48 (Tuesday to Friday) will be given in the lecture room Store Auditorium in Kristen Nygaards hus (IFI1), found here. Student presentations on Monday December 9 will take place in lecture room Prolog in OJDs hus.
Contacts
Jon K. Lærdahl (Course coordinator) - e-mail: jonkl@medisin.uio.no, phone: +47 99 507 335
Torill Rørtveit (Course administrator, registration) - e-mail: torill.rortveit@ibv.uio.no
Computers/laptops, internet access, and UiO user account
All students must bring a laptop with either a Windows (Windows XP or more recent), Unix/Linux, or OS X (i.e. an Apple computer) operating system.
- The computer should not be more than 2-3 years old
- It should be possible to connect the computer to the UiO wireless network
- You must have a root/administrator password that gives you access to installing new software on the computer
- Bring an external mouse, and do not rely on touchpad/trackpad only
- You must have a valid UiO user account and must be able to log onto a computer on the UiO network
- If you are unsure if you have a UiO user account and a valid password, you should try to log in using kiosk.uio.no or win.uio.no as described here. If you are unable to log in, try the hints you find here.
- Instructions (in Norwegian) about how to find your user name and get a new password can be found here.
If you are struggling with anything of the above, in particular if you have forgotten your UiO user name/password or you do not have one, you must contact Jon (See contact details above) as soon as possible, and at least one week before the start of the course.
To get a UiO username/password at the UiO helpdesk you need valid ID.
On the first day of the course we will set up your laptop so that it can be used for the exercises/tutorials, the home exam and hopefully in your future work. How to get a reasonable setup is described here.
If you already are an expert programmer and Unix guru, go here.
Programme
The schedule below is tentative, and may be changed prior to, and possibly even during, the course. Requests and suggestions are welcome.
| Week 46: Monday, November 11 - Friday, November 15 | ||||
| Session 1 | Session 2 | Session 3 | ||
| 09:00 - 10:45 | 11:00 - 12:45 | 13:30 - 16:00 | ||
| Monday 11th |
Course Introduction Biological databases on the web |
Basic Unix tutorial with practicals A tiny taste of simple scripting | ||
| Jon K Lærdahl | Jon K Lærdahl | Jon K Lærdahl | ||
| Tuesday 12th |
Databases/Unix |
Databases/Unix | Databases/Unix | |
| Jon K Lærdahl | Jon K Lærdahl | Jon K Lærdahl | ||
| Wednesday 13th |
Basic Python programming |
Python workshop |
Python workshop | |
| Karin Lagesen | Karin Lagesen | Karin Lagesen | ||
| Thursday 14th |
More Python programming |
Python workshop |
Python workshop | |
| Karin Lagesen | Karin Lagesen | Karin Lagesen | ||
| Friday 15th |
Study day/Start on obligatory take-home assignment | |||
| Week 47: Monday, November 18 - Friday, November 22 | ||||
| Session 1 | Session 2 | Session 3 | ||
| 09:00 - 10:45 | 11:00 - 12:45 | 13:30 - 16:00 | ||
| Monday 18th |
TBA |
TBA |
TBA | |
| TBA | TBA | TBA | ||
| Tuesday 19th |
TBA |
TBA |
TBA | |
| TBA | TBA | TBA | ||
| Wednesday 20th |
TBA |
Chromatin 3D |
Docking and drug discovery - lecture | |
| TBA | Jonas Paulsen | Bjørn Dalhus | Bjørn | |
| Thursday 21st |
TBA |
TBA |
TBA | |
| TBA | TBA | TBA | ||
| Friday 22nd |
Study day/Work on obligatory take-home assignment | |||
| Week 48: Monday, November 25 - Friday, November 29 | ||||
| Session 1 | Session 2 | Session 3 | ||
| 09:00 - 10:45 | 11:00 - 12:45 | 13:30 - 16:00 | ||
| Monday 25th |
Study day/Work on obligatory take-home assignment | |||
| Tuesday 26th |
Introduction to statistical inference and multiple hypothesis testing - lectures |
Basic R programming and exploring your data - lectures/exercises |
Basic statistical testing in R - lectures/exercises | |
| Anja Bråthen Kristoffersen | Anja Bråthen Kristoffersen | Anja Bråthen Kristoffersen | ||
| Wednesday 27th |
Analysis of gene expression data using R - lectures/exercises |
Analysis of gene expression data using R - lectures/exercises |
Analysis of gene expression data using R - lectures/exercises | |
| Ståle Nygård (Anja) | Ståle Nygård (Anja) | Ståle Nygård (Anja) | ||
| Thursday 28th |
Next generation sequencing (NGS) - lectures |
NGS & variant calling lab - lectures/exercises |
NGS & variant calling lab - lectures/exercises | |
| Robert Lyle | Tim Hughes & Robert Lyle | Tim Hughes & Robert Lyle | ||
| Friday 29th |
The bioinformatics of sequencing and assembling genomes - with a focus on the Atlantic cod and salmon genome projects - lectures |
Gene lists & over-representation analysis (ORA) - lectures/practicals |
End of course summary - lectures/discussion | |
| Lex Nederbragt | Ståle Nygård | Jon K Lærdahl | ||
Associate Professor Anja Bråthen Kristoffersen will be present during the exercises/lectures on November 13, 14, and 27.
PhD student Ksenia Khelik from the Department of Informatics will help during some exercises.
On Wednesday November 20, between 11:15 and 12:00, there will be a combined course lecture and CLS seminar by Jonas Paulsen from Oslo University Hospital. This will be on the study of chromatin 3D structure.
Exam
The exam for this course will be a one week, take-home exam.
The exam will be sent to all participants at 3 pm, Thursday December 6, by e-mail.
Your completed exam must be returned, at the latest, at 3 pm, Friday December 14. It should be sent by e-mail to the course administrator Torill Rørtveit (e-mail address: torill.rortveit@ibv.uio.no). Please put the course code and your candidate number in the subject field (e.g. "Exam MBV-INF4410 Candidate:12345").
The exam must be handed in as a single PDF document (Microsoft Word or an Open Office Document is also acceptable). The document should be named with the course code and your candidate number only (e.g. MBV-INF4410-12345.pdf). Do not place your name in the document.
Written assignment
Stuff
Bioinformatics mailing list for the Oslo region
The mailing list for computational biology and bioinformatics in the Oslo region is cbo-all@usit.uio.no. The list has approximately 370 members. The list is used to distribute news about seminars, positions, courses, meetings and other topics that might be of interest to students and researchers with an interest in computational life science in south-eastern Norway. If you want to receive e-mails that are sent to the list, sign up here
https://sympa.uio.no/usit.uio.no/info/cbo-all
by following the link termed "Subscribe".
Useful links
Trond Hasle Amundsen's Local guide to Linux and Unix
EMBnet Quick guide Unix
UCSC Genome browser
Free UCSC Genome browser tutorial from OpenHelix
Portal to Galaxy
Galaxy 101 and other Galaxy screencasts/tutorials
UiO Lifeportal and more information on the Lifeportal
The Genomic HyperBrowser