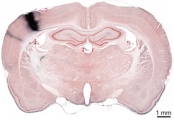
Whole Brain Connectivity Atlas
Whole Brain Connectivity Atlas is part of the Rodent Brain WorkBench, a new research and development project funded by The Research Council of Norway, the Centre for Molecular Biology and Neuroscience, and the European Union.
Contents
About
The Whole Brain Connectivity Atlas is an interactive resource providing access to brainwide experimental tract-tracing data showing the distribution of axonal connections arising from the primary somatosensory cortex (SI) in the rat brain.
The atlas is a resource for mapping and sharing of connectivity data derived from histological axonal tract-tracing experiments. It has been developed to provide access to high-resolution images of histological sections showing the distribution of anterogradely labelled axons arising from the rat primary somatosensory and primary motor cortex (SI ans MI).
Currently, the atlas contains the results from six experiments in which an anterograde tracer, either biotinylated dextran amine (BDA) or Phaseolus vulgaris leucoagglutinin (Pha-L), was injected in the SI forelimb or whisker representations in M1. The results include images of ~1500 coronal sections showing the distribution of tracer labelled axons across the entire rat brain.
- SIwiring.jpg
Cortical and subcortical projections of SI (modified from Welker et al., Exp Brain Res 73:411-435).
Features
- Tract-tracing results from six experiments in which an anterograde tracer was injected in the forelimb or whisker representations in the rat SI or MI
- About 1500 images of coronal sections showing:
- the distribution of labelling (BDA or Pha-L)
- cytoarchitectural features (Neutral red or thionine counterstaining)
- SI chemoarchitecture (every other section stained for cytochrome oxidase)
- assigned bregma values based on the anatomical features revealed by the NR, CO and thionine staining, corresponding to a standard stereotaxic space of the rat brain (Paxinos & Watson, 2007)
Methods
The tract-tracing experiments were conducted in four adult Sprague Dawley rats and two adult Wistar rats. In each animal, an anterograde axonal tracer (biotinylated dextran amine, BDA or Phaseolus vulgaris leucoagglutinin, Pha-L) was iontophoretically injected in the whisker or forelimb representation of SI (five cases) or MI (one case). Coronal brain sections were stained to visualize BDA or Pha-L and counterstained to visualize cytoarchitectural features. Selected sections were stained for cytochrome oxidase to visualize the characteristic SI barrel organization. Images were obtained through an automated Olympus Bx52 microscope, equipped with a high-precision motorized stage running the Neurolucida v6.0 Virtual Slice software (MicroBrightField Inc., Williston, VT, USA). (For further details, see Zakiewicz et al., 2011, PLoS One, 6(8):e22669)
- NR vs no stain final.jpg
The location of injection sites and BDA labelling can be localized on the basis of anatomical landmarks and cytoarchitectonic features.
- Injectionsites final.jpg
Weak counterstaining with Neutral Red is used to visualize cytoarchitecture without compromising the pattern of BDA labelling.
Staining abbreviations
|
Abbreviations |
Description |
|
CO |
Stained for cytochrome oxidase |
|
BDA_ABC |
Stained for biotinylated dextran amine, ABC method |
|
BDA_ABC_NR |
Stained for biotinylated dextran amine, ABC method, counterstained with Neutral Red |
|
Pha-L_PAP |
Stained for Phaseolus vulgaris leucoagglutinin, PAP method |
|
Pha-L_PAP_thio |
Stained for Phaseolus vulgaris leucoagglutinin, PAP method, counterstained with thionine |
|
Pha-L_ABC |
Stained for Phaseolus vulgaris leucoagglutinin, ABC method |
|
Pha-L_ABC_thio |
Stained for Phaseolus vulgaris leucoagglutinin, ABC method, counterstained with thionine |
News
2015-04 Atlas moved to NeSys wiki. New image viewer was made available and additional image material has been added
2014-02 Second article with examples of analysis of the rat SI connectivity based on the histological data from the existing database was published: [1]
2011-08 Open access following publication in PLoS ONE
2011-07 First article published, including an open source platform with histological data from six axonal tract-tracing experiments: [2]
2009-09 Poster / demonstration, 2nd INCF Congress of Neuroinformatics, Plzen, Czech Republic
Access images
Re-use of data from this repository is allowed provided that reference is given to the following publication: Zakiewicz IM, van Dongen YC, Leergaard TB, Bjaalie JG (2010) Workflow and database system for mapping the axonal connectivity of the rat brain. PLoS ONE 2011;6(8):e22669
The image repository contains ~1200 coronal images showing data from 6 axonal tracing experiments. The viewer allows interactive zooming and panning. Select experiment of interest in the list below. For further details, see Tutorial
| Injection site |
Location area | Images | About the case |
|---|---|---|---|
| |
SI forelimb, BDA (R601) | Open viewer | More details (pdf) |
| |
SI whisker D2, BDA (R602) | Open viewer | More details (pdf) |
| |
SI forelimb, BDA (R603) | Open viewer | More details (pdf) |
| |
SI whisker D3, BDA (R604) | Open viewer | More details (pdf) |
| |
forelimb, Pha-L (R605) | Open viewer | More details (pdf) |
| |
SI whisker D5, Pha-L (R606) | Open viewer | More details (pdf) |
Tutorial
To use the Image repositories and viewer, first drag the slider below the row of thumbnail images horizontally to select an anterioposterior level of interest (bregma coordinates are indicated). Click on a thumbnail image to open it in the viewer.
The viewer has the following options for navigating the images:
- 'Zoom' (zoom in and out by using mouse scroll button or two-finger pan on touchpad)
- 'Pan' (left-click and drag the images freely in the viewer, after enlarging the image you can chose the area of interest by left-clicking on the preview window)
- 'Reset' (click “view all” button)
- 'Annotation (under development)
The information about selected section is avaliable in the upper right corner with the possibility for downloading the image. To download images (at viewer resolution) use the download button in the upper right corner. To print images, either use the print screen functionality of your computer.
Each section has assigned bregma value, visible under every thumbnail and in the image information corner. The value was assigned based on section landmarks revealed by counterstain or CO-staining and corresponds to the popular stereotaxic rat brain atlas (The Rat Brain In Stereotaxic Coordinates, Paxinos and Watson, 2007).
Work in progress
- Preparation of interactive tabular listing of label distributions, reciprocally linked with image viewer
- 3D reconstruction of projection patterns (article submitted for reviewing)
- 3D data / connectivity project (co-registration within Waxholm space)
Documentation
Connectivity atlas system:
Zakiewicz IM, van Dongen YC, Leergaard TB, Bjaalie JG (2010) Workflow and database system for mapping the axonal connectivity of the rat brain. Submitted manuscript
Zakiewicz IM, van Dongen YC, Moene IA, Leergaard TB and Bjaalie JG (2009) Towards an interactive digital atlas system for exploring anatomical connectivity patterns in the entire rat brain. Frontiers in Neuroinformatics. Conference Abstract: 2nd INCF Congress of Neuroinformatics. doi: 10.3389/conf.neuro.11.2009.08.014
Database architecture:
Moene IA, Subramaniam S, Darine D, Leergaard TB, Bjaalie JG (2007). Towards a workbench for rodent brain image data: systems architecture and design. Neuroinformatics; 5: 35-58
Bjaalie JG, Leergaard TB, Lillehaug S, Odeh F, Moene IA, Kjøde JO, Darine D (2005) Database and tools for analysis of topographic organization and map transformations in major projection systems of the brain. Neuroscience 2005; 136:681-696
Contact
Connectivity and digital brain atlasing: j.g.bjaalie@medisin.uio.no
Technical issues and functionality: d.a.darine@medisin.uio.no
Credits
NeSys and CMBN atlas development team:
Izabela M. Zakiewicz, M.Sc.
Yvette C. van Dongen, Ph.D
Anna Torbjørg Bore
Jan Olav Kjode, M.Sc.
Ivar Andre Moene, M.Sc.
Trygve B. Leergaard, M.D., Ph.D.
Jan G. Bjaalie, M.D., Ph.D.
Funded by The Research Council of Norway, the Centre for Molecular Biology and Neuroscience, and the European Union
Contributing laboratories
Neural Systems and Graphics Computing Laboratory
Centre for Molecular Biology and Neuroscience & Institute of Basic Medical Sciences
Deptartment of Anatomy
University of Oslo
P.O. Box 1105 Blindern
N - 0317 Oslo
Norway
http://www.nesys.uio.no