WRFand WRF-CHEM
An example of running WRF can be found here: wiki.uio.no/mn/geo/geoit/index.php/A_WRF_example.
Contents
Weather Research and Forecasting Model
WRF and WRF-CHEM have been installed on Abel. To check which version is available:
module avail wrf -------------------------------------------------------------------------------------- /cluster/etc/modulefiles --------------------------------------------------------------------------------------- wrf/3.6.1
To set-up your environment:
module load wrf/3.6.1
When loading wrf, the following environment variables are defined:
- WRF_HOME: directory where WRF has been installed; this can be used to check WRF sources
- WPS_HOME: directory where WPS has been installed. It is unlikely you have to check or change WPS source code but you will find in this directory all the metadata required to run WPS (the GEOGRID.TBL, METGRID.TBL, and Vtable files).
- WPS_GEOG_PATH: contains theterrestrial static input data. Even if you use your own compiled version of WRF/WPS, it is NOT necessary to download these files again.
- WRFIO_NCD_LARGE_FILE_SUPPORT: it is set to 1 to allow you to store large netCDF files (more than 2GB).
- WRF_EXAMPLES: directory where all WRF and WRF-CHEM examples are stored. If you wish to run one of this example see our dedicated section on running tutorials.
WRF has been compiled with intel compilers and MPI:
module list Currently Loaded Modulefiles: 1) use.own 3) openmpi.intel/1.6.1 5) netcdf.intel/4.2.1.1 7) jasper/1.900.1 9) openmpi.intel/1.8 11) wrf/3.6.1 2) intel/2011.10 4) hdf5/1.8.9_intel 6) intel-libs/2013.sp1.3 8) intel/2013.sp1.3 10) ncl/6.2.0
We suggest you specify the version you wish to use to avoid any problems if we install a new default version (as it is important to stick to the very same version for your simulations).
Running WRF:
The following steps are necessary to run WRF on our systems:
- WRF Preprocessing System (WPS)
- WRF System
WRF Preprocessing System (WPS)
Most parameters for WPS must be given in namelist.wps
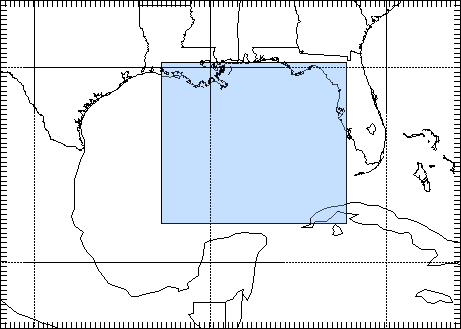
- define_grid.py uses namelist.wps to plot your defined grid
- run_geogrid.py runs geogrid.exe to create static data for your domain
- run_ungrib.py runs ungrib.exe to unpack your input GRIB data
- run_metgrid.py runs metgrid.exe to interpolate input data onto your model domain
WRF
Most parameters for WRF must be given in namelist.input
- run_init_wrf.py runs real.exe to initialize WRF and creates two files such as wrfinput_d<domain> and wrfbdy_d<domain> where <domain> is the domain number (01, 02, etc.)
- run_wrf.py runs the numerical integration program wrf.exe
TIPS: to get the usage of these python scripts, use -h or --help. For instance:
run_ungrib.py -h
Usage: run_ungrib.py --expid expid --vtable vtable --datadir datadir [--start_date startdate --end_date enddate] [--interval_seconds val] [--prefix FILE]
Options:
-h, --help show this help message and exit
-s startdate, --start_date=startdate
A list of MAX_DOM character strings of the form
'YYYY-MM-DD_HH:mm:ss' specifying the starting UTC date
of the simulation for each domain.
-e enddate, --end_date=enddate
A list of MAX_DOM character strings of the form 'YYYY-
MM-DD_HH:mm:ss' specifying the ending UTC date of the
simulation for each domain
-t interval_seconds, --interval_seconds=interval_seconds
The integer number of seconds between time-varying
meteorological input files. No default value.
-p FILE, --prefix=FILE
prefix of the intermediate meteorological data files
-v Vtable, --vtable=Vtable
vtable filename
-i expid, --expid=expid
Experiment identifier
-d datadir, --datadir=datadir
Directory where input fields can be found
For each of these python scripts, some arguments are optionals and indicated in squared brackets such as start_date and end_date. If you don't specify them on the command line, it will keep what has been defined in your namelist. This is usually how you will run your "real" cases.
WRF examples:
All WRF examples can be found in $WRF_EXAMPLES
A subdirectory can be found for each example and it contains all you need to run the corresponding examples (namelist.wps, namelist.input, Vtable, workflow.bash, and all the DATA required to run the simulation). You may found one or more files named workflow*.bash. These files are describing the sequence of programs to run for each example and can be divided in two groups:
January2000Case
This case is the East Coast Winter Storm of January 24-25, 2000 (http://cimss.ssec.wisc.edu/goes/misc/000125.html)
To run it, you need to execute each of the statement in workflow.bash:
> cat workflow.bash
#!/bin/bash
# check your domain is OK
define_grid.py --path /cluster/software/VERSIONS/wrf/examples/January2000Case
# Run geogrid.exe to create static data for this domain:
run_geogrid.py -p /cluster/software/VERSIONS/wrf/examples/January2000Case --expid January2000Case
# Unpack input GRIB data (ungrib.exe)
run_ungrib.py --expid January2000Case \
--start_date 2000-01-24_12:00:00 \
--end_date 2000-01-25_12:00:00 \
--interval_seconds 21600 \
--prefix FILE \
--vtable /cluster/software/VERSIONS/wrf/examples/January2000Case/Vtable \
--datadir /cluster/software/VERSIONS/wrf/examples/January2000Case/DATA/
#Interpolate the input data onto our model domain (metgrid.exe)
run_metgrid.py --expid January2000Case
# Initialize WRF model (real.exe/ideal.exe)
run_init_wrf.py --expid January2000Case \
--namelist /cluster/software/VERSIONS/wrf/examples/January2000Case/namelist.input
# Run the model (wrf.exe)
run_wrf.py --expid January2000Case \
--namelist /cluster/software/VERSIONS/wrf/examples/January2000Case/namelist.input
You don't have to change paths or namelists. it has been set-up to execute WPS/WRF in your workdir ($WORKDIR) and a subdirectory named January2000Case (named from your experiment identifier given with --expid option).
To visualize your outputs (named wrfout*), you may use ncview:
module load ncview
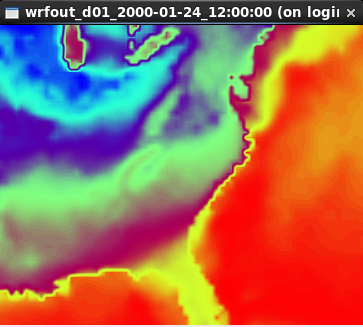
ncview wrfout_d01_2000-01-24_12:00:00
and select the variables you wish to plot. For instance, to visualize SST:
| |
|
TIPS: ncview is not recommended for quality graphical displays, but is a very handy tool for a quick first-look at the data.
HurricaneKatrina
As for previous examples, you just need to run workflow.bash and check the directory $WORKDIR/HurricaneKatrina
On August 28, 2005, Hurricane Katrina was in the Gulf of Mexico, where it strengthened to a Category 5 storm on the Saffir-Simpson hurricane scale, packing winds estimated at 175 mph. (http://www.katrina.noaa.gov/).
| Hurricane Katrina on August 28, 2005 (image taken from http://www2.mmm.ucar.edu/wrf/OnLineTutorial/CASES/SingleDomain/) |
HurricaneKatrinaSST
The goal here is to input SST into WRF model. For these runs we will use the Hurricane Katrina case data (2005-08-28_00 to 2005-08-29_00).
SST are typically added to the model:
a. Use the SST at the initial time as a constant field for all time periods (this is good for short runs, like real-time runs, where SST is not updated during the WRF model run)
b. As an extra input at each model input time (this is good for long -months- model runs)
NestedModelRuns
For these runs we will use the Katrina Hurricane case data (2005-08-28_00 to 2005-08-29_00).
The domain we are going to set up is show below (image taken from http://www2.mmm.ucar.edu/wrf/OnLineTutorial/CASES/NestRuns/index.html).
| |
There are a number of different ways to set up nested model runs (in this tutorial we are only going to set-up 2-way interactive nested runs).
a. Two-way nested run, with one input file
The preprocessing steps for this case will be similar to a single domain setup. The only difference, is that during the wrf.exe execution, a second (or more) nest(s) is initiated. The corresponding workflow can be found in workflow_a.bash
b. Two-way nested run, with two input files
For this case the pre-processing programs need to be run to create extra input data for the wrf model run. At the WRF model step, one has the choice to:i. Use all the meteorological and static data for nested domains as input, (see workflow_b.bash) or
ii. Use only the static data for nested domains as input (see workflow_bb.bash).c. One-way nesting using ndown
ndown is used to run one-way nested runs AFTER wrf has already been run for the mother domain.
One-way nesting can also be done similar to two-way nested runs (both a and b above), by simply setting feedback in the WRF namelist.input file equal to 0. The corresponding workflow can be found in workflow_c.bash
RestartRun
This case study we will use the same setup as for the Single Domain run, we will just restart it from the previous run. As for all other examples, run workflow.bash
April2005Case
This example is showing how to run from ERA-Interim data instead of GFS data. As you can see in workflow.bash, another Vtable needs to be used (Vtable.EI).
WRF has been compiled with WRF_CHEM=1 and WRF_KPP=1 and is therefore suitable for WRF-CHEM simulations. The three next examples shows how to use WRF-CHEM at UIO.
BiogenicEmissions
This example uses WRF-CHEMand its goal is to get familiar with the methodology by which the MEGAN biogenic emissions are introduced into the WRF-Chem simulation.
This exercise is intended to be completed by students that have knowledge about setting-up and running the WRF numerical model.
There are two workflows:
- option-1 (workflow-opt1.bash) uses GOCART-RACM_KPP aerosol option (chem_opt=301), Geunther biogenic emissions (bio_emiss_opt=1). dust, sea salt, DMS, and biomass burning will still be included so keep those options turned on.
- option-2 (workflow-opt2.bash)
DustErosion2010
This example uses WRF-CHEM. The purpose of this example is to get familiar with the methodology by which the dust erosion fields are introduced through the WRF Preprocessing System (WPS). The corresponding workflow is calledworkflow.bash
GOCARTaerosols
This example uses WRF-CHEM. A global emissions data set was prepared by a program called "prep_chem_sources" and with this program anthropogenic emissions, GOCART background fields and biomass burning (wild fire) emissions was previously mapped to the user domain. In this exercise you will use the emissions data and follow the methodology for making a WRF-Chem forecast shown here. The corresponding workflow is called workflow.bash
Running "long" simulations on abel
SLURM batch system
For most of your WRF runs (wrf.exe), you will need to use SLURM batch system (you can not use more than 30mn CPU on the interactive node and cannot run WRF efficiently in parallel). This requires to create what we call a batch job script: it's a script with additional SLURM directives. SLURM is the current batch system used on abel.
The most important when running wrf is to:
- choose the number of tasks (#SBATCH --ntasks)
- set the amount of memory per task (#SBATCH --mem-per-cpu)
- set the wall clock time limit (#SBATCH --time)
and add the following SLURM directives in your script:
# Number of tasks (cores): #SBATCH --ntasks=8
it sets the number of tasks to 8 (i.e. WRF will be using 8 processors for running wrf.exe)
# Max memory usage per task: #SBATCH --mem-per-cpu=4000M #
it sets the maximum amount of memory you will be using per tasks. In the above example, 4000 Mb per task may be used.
# Wall clock limit: #SBATCH --time=100:0:0
it sets a limit of 100 hours for your run. It is advised to split long simulations in chunks and create restart files regularly instead of submitting a huge job.
For more information on the queue system, look here.
Save your outputs on your local machine
When running WRF, your outputs will be stored in $WORKDIR and deleted after about 45 days. It is therefore important to save your outputs on your local machine. This is usually done with an rsync command (see example below) and to avoid having to enter your password everytime you invoke this command, you must set your SSH-keys:
% cd ~/.ssh
% ssh-keygen -t rsa
Generating public/private rsa key pair.
Enter file in which to save the key (~/.ssh/id_rsa): (just type return or something like ~/.ssh/id_rsa_name of my machine)
Enter passphrase (empty for no passphrase): (just type return)
Your identification has been saved in ~/.ssh/id_rsa
Your public key has been saved in ~/.ssh/id_rsa.pub
The key fingerprint is:
Some really long string
Step 2:
Then, paste content of the local ~/.ssh/id_rsa.pub file into the file ~/.ssh/authorized_keys on the remote host. Please make sure the file ~/.ssh/authorized_keys is readbale by you only:
chmod og-rwx ~/.ssh/authorized_keys
You can do Step 1 on your local machine and Step 2 on abel and then Step 1 on abel and Step 2 on your local machine to have access from and to abel.
Once this is done, you can use rsync command or scp to copy your data back to your local directory. An example in given in the complete batch script given below.
Example of a complete batch script for running wrf on abel
Here we run wrf.exe for the January200Case (see WRF examples). We assume that all the previous steps have been run and run wrf.exe in batch mode.
#!/bin/bash
# Job name:
#SBATCH --job-name=wrf
#
# Project:
#SBATCH --account=geofag
#
# Wall clock limit:
#SBATCH --time=100:0:0
#
# Max memory usage per task:
#SBATCH --mem-per-cpu=4000M
#
# Number of tasks (cores):
#SBATCH --ntasks=8
#
# Set up job environment (do not remove this line...)
source /cluster/bin/jobsetup
ulimit -s unlimited
module load wrf
# Run the model (wrf.exe)
run_wrf.py --expid January2000Case --npes 8 \
--namelist /cluster/software/VERSIONS/wrf/examples/January2000Case/namelist.input
# Copy your data on your local machine
rsync -avz $WORKDIR/January2000Case/wrfout* $USER@sverdrup.uio.no:January2000Case/.
rsync -avz $WORKDIR/January2000Case/wrfrst* $USER@sverdrup.uio.no:January2000Case/.
#
# End of jobscript
#
To submit your job script:
sbatch job_wrf.sh
For more information on Abel see The Abel compute Cluster.
How to change or add new code?
The easiest for changing WRF code is to copy the pre-installed version, make your changes and then recompile WRF. Here is an example:
module load wrf mkdir $HOME/WORK cd $HOME/WORK cp -R $WRF_HOME . module purge export WRF_HOME=$HOME/WORK/WRFV3 module load wrf
Then you can edit any files and recompile WRF:
./clean -a
./configure
(make sure you choose 15 i.e. dmpar)
./compile em_real >& compile.log
It is unlikely you have to change WPS but if you need to, you should do the same (define WPS_HOME, etc.).
TIPS and known problems
Often-seen runtime problems
- Segmentation fault (core dumped): if it appears immediately after starting wrf.exe, it may be due to insufficient stack memory. Try:
ulimit -s unlimited
and rerun wrf.exe
Problems for creating all the WRF inputs when having more than one domain (and get wrfinput_d01 and wrfbdy_d01 only)
- if your namelist.wps has the correct list of dates for each domain, make sure you do not use start_date and end_date option when running run_ungrib.py
For instance:
run_ungrib.py --expid NestedModelRunsB \
--vtable /cluster/software/VERSIONS/wrf/examples/HurricaneKatrina/Vtable.GFS \
--datadir /cluster/software/VERSIONS/wrf/examples/HurricaneKatrina/DATA/Katrina
- If you wish to pass the list of dates, make sure you give the dates for each domain:
run_ungrib.py --expid NestedModelRunsB \
--start_date "2005-08-28_00:00:00, 2005-08-28_00:00:00" \
--end_date "2005-08-29_00:00:00, 2005-08-28_00:00:00" \
--interval_seconds 21600 \
--prefix FILE \
--vtable /cluster/software/VERSIONS/wrf/examples/HurricaneKatrina/Vtable.GFS \
--datadir /cluster/software/VERSIONS/wrf/examples/HurricaneKatrina/DATA/Katrina
Possible error messages if the data download is incorrect
Landsea mask
When running metgrid, the following error may result if you download constant fields for the surface:
/usit/abel/u1/irenebn/Scandinavia/April2005Case/METGRID.TBL Processing domain 1 of 1 LSM:2014-04-15_12 Z:2014-04-15_12 Processing 2014-04-15_00 FILE ERROR: Cannot combine time-independent data with time-dependent data for field LANDSEA.mask -------------------------------------------------------------------------- MPI_ABORT was invoked on rank 0 in communicator MPI_COMM_WORLD with errorcode 0. NOTE: invoking MPI_ABORT causes Open MPI to kill all MPI processes. You may or may not see output from other processes, depending on exactly when Open MPI kills them. --------------------------------------------------------------------------
This error resulted from having downloaded lsm (variable 129.128), and is solved by avoid downloading constant fields.