Software and resources
Contents
Software and resources
Overview of codes from the Materials Design Group at Imperial College London
Installation on Saga
module load Python/3.9.6-GCCcore-11.2.0 pip install sumo pip install --upgrade https://github.com/SUNCAT-Center/catmap/zipball/master
This will install the python packages under .local/lib/python3.9/site-packages/
Structure and crystallography
ICSD database
Automatic login through EZproxy (www.ub.uio.no/english/using/remote-access.html).
Materials project
Contains structures optimized by DFT that can be downloaded in several formats including .cif or POSCAR. Choose 'Conventional Standard' CIF.
Login with a Google account (available with UiO username www.uio.no/english/services/it/store-collaborate/gsuite/).
International Tables for Crystallography
Complete overview of space-group symmetry and more.
Shannon Ionic Radii
Reference [1]
Binding energies of molecules
Todo: Table of binding energies with reference (e.g., NIST)
Catalysis
Catalysis Hub
Database of reaction energies and barriers from DFT calculations.
No entropy found
Visualization
VESTA
jp-minerals.org/vesta/en/ (Windows, macOS, Linux)
View periodic structures, charge densities and more.
Recommended settings for improved figures
Objects > Properties > Atoms/Bonds/Polyhedra Specular: 40 40 40 Shininess (%): 1 View > Overall Appearance... Ambient: 10 Diffuse: 70
Avogadro
avogadro.cc (Windows, macOS, Linux)
View and edit molecular structures and optimize molecular geometry through molecular mechanics.
Diamond
www.crystalimpact.com/diamond/ (Windows: Licence)
View and edit periodic and molecular structures.
VMD
www.ks.uiuc.edu/Research/vmd/ (Windows, macOS, Linux)
View and animate structures from molecular dynamics simulations.
P4vasp
github.com/orest-d/p4vasp (macOS, Linux)
Visualizing periodic structures, density of states and band structures.
OVITO
OVITO is a visualization and analysis software for output data generated in molecular dynamics, atomistic Monte-Carlo and other particle-based simulations.
Escher
http://schooner.chem.dal.ca/wiki/Escher
Escher is a collection of octave and python routines for the visualization and plotting of molecules and crystals. The original idea in escher was to modernize tessel, a program for the representation of crystal structures (and more). Via an interface to POV-Ray, tessel is able to generate very high quality representations of periodic and finite molecular structures.
Code link of a cover art example: https://github.com/heesoopark/Escher-PovRay
Packages
Spyder
www.spyder-ide.org (Windows, macOS, Linux)
Scientific python developer environment.
Atomic Simulation Environment (ASE)
Set of tools and Python modules for setting up, manipulating, running, visualizing and analyzing atomistic simulations.
ASE modules are available on Saga
module avail ase
Example of a script for generating an (1 1 1) surface slab of palladium
#!/opt/local/bin/python from ase.io import read from ase.io import write from ase.build import fcc111 from ase.build import fcc100 from ase.build import fcc111_root
slab = fcc111_root('Pd', 3, size=(1,2,7), a=3.9438731474981594, vacuum=5.0)
write('Pd-111.cif', slab, 'cif')
VASPKIT
VASPKIT provides a powerful and user-friendly interface to perform high throughput analysis of various material properties from the raw calculated data using the widely-used VASP code.
- Generate KPOINTS, POTCAR and INCAR for a given POSCAR file;
- Elastic-constants of 2D and bulk materials using stress-strain or energy-strain methods;
- Equation-of-state fitting;
- Suggested k-paths for a given crystal structure;
- Optical adsorption coefficient of 2D and bulk materials;
- Band structure unfolding;
- Fermi surface;
- Density-of-states and band-structure;
- Charge/spin density, Charge density difference;
- Vacuum level and work function;
- Wave-function analysis;
- Molecular-dynamics analysis;
- Effective mass of carrier;
- Symmetry finding and operations;
- 3D band structures;
- Magnetocrystalline anisotropy energy;
- Currently, only VASP raw data are fully supported.
CatMAP
CatMap is a catalyst Micro-kinetic Analysis Package for automated creation of micro-kinetic models used for studying kinetics and screening new catalysts.
https://catmap.readthedocs.io/en/latest/index.html
CatMap can be installed directly via pip:
pip install --upgrade https://github.com/SUNCAT-Center/catmap/zipball/master
You may need to add the install directory to the PYTHONPATH, e.g. in bash shell or .bash_profile
export PYTHONPATH=$HOME/THIS_FOLDER_PATH:$PYTHONPATH
Input File Structure
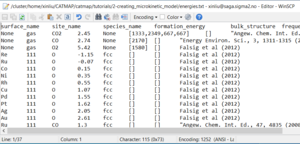
The TableParser accepts inputs in a tab-separated text file. An example of the header and first few lines are provided below:
The following column titles are required for a functional input file:
- surface_name
- site_name
- species_name
- formation_energy
- frequencies
- reference
Generating input file for formation energy
You can use the python script under /CATMAP/catmap/tutorials/1-generating_input_file/generate_input.py
Creating a Microkinetic Model
Check the example of CO oxidation in the path of /CATMAP/catmap/tutorials/2-creating_microkinetic_model
One of the most important aspects of the “setup file” is the “rxn_expressions” variable which defines the elementary steps in the model. For this simplified CO oxidation model we will specify these as:
rxn_expressions = [ '*_s + CO_g -> CO*', '2*_s + O2_g <-> O-O* + *_s -> 2O*', 'CO* + O* <-> O-CO* + * -> CO2_g + 2*', ]
The first expression includes CO adsorption without any activation barrier. The second includes an activated dissociative chemisorption of the oxygen molecule, and the final is an activated associative desorption of CO2. More complex models for CO oxidation could be imagined, but these elementary steps capture the key features. Note that we have only included “*” and “*_s” sites since this is a single-site model for CO oxidation. This means that all intermediates will be adsorbed at a site designated as “s”. These reaction expressions will be parsed automatically in order to define the adsorbates, transition-states, gasses, and surface sites in the model.
Phonopy and Phono3py
Phonopy is an open source package for phonon calculations at harmonic and quasi-harmonic levels. Phono3py is another open source package for phonon-phonon interaction and lattice thermal conductivity calculations.
Features
- Phonon band structure, phonon DOS and partial-DOS
- Phonon thermal properties: Free energy, heat capacity (Cv), and entropy
- Phonon group velocity
- Thermal ellipsoids / Mean square displacements
- Irreducible representations of normal modes
- Dynamic structure factor for INS and IXS
- Non-analytical-term correction: LO-TO splitting (Born effective charges and dielectric constant are required.)
- Mode Grüneisen parameters
- Quasi-harmonic approximation: Thermal expansion, heat capacity at constant pressure (Cp)
- Interfaces to calculators: VASP, VASP DFPT, ABINIT, Quantu ESPRESSO, SIESTA, Elk, WIEN2k, CRYSTAL, DFTB+, TURBOMOLE, CP2K, FHI-aims, CASTEP, Fleur, LAMMPS (external)
- Phonopy API for Python
TDEP
Extract force constants, phonon dispersion relations, thermal conductivity, and generate special quasirandom structures (SQS)
Available on Saga (/cluster/shared/tdep/bin). Use the following modules
module purge module load Anaconda3/2019.03 module load intel/2018b module load imkl/2018.3.222-iimpi-2018b module load HDF5/1.10.2-intel-2018b
Generate SQS supercell from from a unit cell POSCAR file named 'infile.ucposcar'
Example of 'infile.ucposcar' for a A-site doped SrTiO3 unit cell where the disordered site is designated 'ALLOY' (2 elements: 52% Sr and 48% Ca)
Sr1 Ti1 O3 1.0 3.945130 0.000000 0.000000 0.000000 3.945130 0.000000 0.000000 0.000000 3.945130 ALLOY Ti O 1 1 3 direct 0.000000 0.000000 0.000000 2 Sr 0.52 Ca 0.48 0.500000 0.500000 0.500000 0.500000 0.000000 0.500000 0.500000 0.500000 0.000000 0.000000 0.500000 0.500000
Generate 2x2x2 SQS supercells (five supercells will be generated outfile.sqs_001-005)
generate_structure -d 2 2 2
Supercell
https://orex.github.io/supercell/
Supercell is a combinatorial structure-generation approach for the local-level modeling of atomic substitutions and partial occupancies in crystals.
Supercell can generate special quasirandom structures (SQS).
Icet
Icet is a module in Python that can generate special quasirandom structures. [1]
Here is an example of a script to generate an SQS structure of a 3x3x3 supercell of a rocksalt structure ( TiN ) with several H atoms on the N sites (TiN0.75H0.25)
from ase import Atom
from ase.build import bulk
from icet import ClusterSpace
from icet.tools.structure_generation import (generate_sqs,
generate_sqs_from_supercells,
generate_sqs_by_enumeration,
generate_target_structure)
from ase.io import write
#makes a TiN cell
primitive_structure = bulk('TiN', cubic=True, crystalstructure='rocksalt', a=4.2570)
#makes a 3x3x3 supercell
supercells = [primitive_structure.repeat(3)]
# defines a cluster space. There are 8 brackets because we have 8 atoms in the cell defined earlier (Ti_4N_4). Several aloms in the same bracket mean that they are allowed on the same site
cs = ClusterSpace(primitive_structure, [7.0,4.5],chemical_symbols=[['Ti'], ['N','H'],['Ti'], ['N','H'],['Ti'], ['N','H'],['Ti'], ['N','H']])
# x in TiNx
x = 0.75
# In this case we want TiN_{0.75}H_{0.25}. We only act on sublattice A because we do not want to change the concentration of Ti. The target concentration has to be compatible with the supercell (i.e. it should correspond to an integer number of atoms in the cell)
target_concentrations = {'A': {'N': x, 'H': 1-x}}
#here we use “generate_sqs_from_supercells” because it allows us to make a supercell that has the same geometry as the one in the input
# With more steps, we generate better structures, it also takes longer
sqs=generate_sqs_from_supercells(cluster_space=cs,
supercells=supercells,
target_concentrations=target_concentrations,
n_steps=50000 )
write('TiNH.cif', sqs)
Spinney
Python package dedicated to the study of point defects in solids. Can be used to calculate the correction energy due to electrostatic finite-size-effects in charged supercells, defect formation energies and transition levels, and defects concentrations.
SUMO
Sumo is a Python toolkit for plotting and analysis of ab initio solid-state calculation data, built on existing Python packages from the solid-state chemistry/physics community. SUMO can plot projected DOS directly from DOSCAR and vasprun.xml
Installation on Saga works with the following python module (consider loading it in ~/.bash_profile)
module load Python/3.9.6-GCCcore-11.2.0
pip install sumo
The installation folder will be ~/.local/lib/python3.9/site-packages/
Pymatgen
Pymatgen (Python Materials Genomics) is a robust, open-source Python library for materials analysis.
Generate a POTCAR with symbols Li_sv O and the PBE functional in a termial
pmg potcar --symbols Li_sv O --functional PBE
Python code quick examples:
1) to generate NEB images by using IDPP method (image dependent pair potential: J. Chem. Phys. 140, 214106 (2014))
from pymatgen.core import Structure
from pymatgen.analysis.diffusion.neb.pathfinder import IDPPSolver
init_struct = Structure.from_file("POSCAR_start.vasp")
final_struct =Structure.from_file("POSCAR_end.vasp")
obj = IDPPSolver.from_endpoints(endpoints=[init_struct, final_struct], nimages=5,
sort_tol=1.0)new_path = obj.run(maxiter=5000,
tol=1e-5, gtol=1e-3, species=["Na"])
for i in range(len(new_path)):
print(new_path[i])
new_path[i].to(fmt="poscar",filename="POSCAR-"+str(i)+".vasp")
2) to analyze NEB results
from pymatgen.analysis.transition_state import NEBAnalysis
neb = NEBAnalysis.from_dir("./")
nebplot = neb.get_plot()
nebplot.savefig("NEB.png")
Atomsk
Atomsk creates, manipulates, and converts data files for atomic-scale simulations in the field of computational materials sciences in CLI. Atomsk can generate common lattice types such as fcc, bcc, hcp, and perform elementary transformations: duplicate, rotate, deform, insert dislocations, merge several systems, create twins, bicrystals and polycrystals, and so on. These elementary tools can be combined to construct and shape a wide variety of atomic systems.
To compile on Saga by using Intel fort/MLK,
git clone https://github.com/pierrehirel/atomsk.git
cd atomsk/src
module load intel-compilers/2022.1.0
module load imkl/2022.1.0
make -f Makefile.ifort atomsk
References
- ↑ R.D. Shannon, Revised Effective Ionic Radii and Systematic Studies of Interatomic Distances in Halides and Chalcogenides, Acta Cryst. 1976 (A32) 751-767